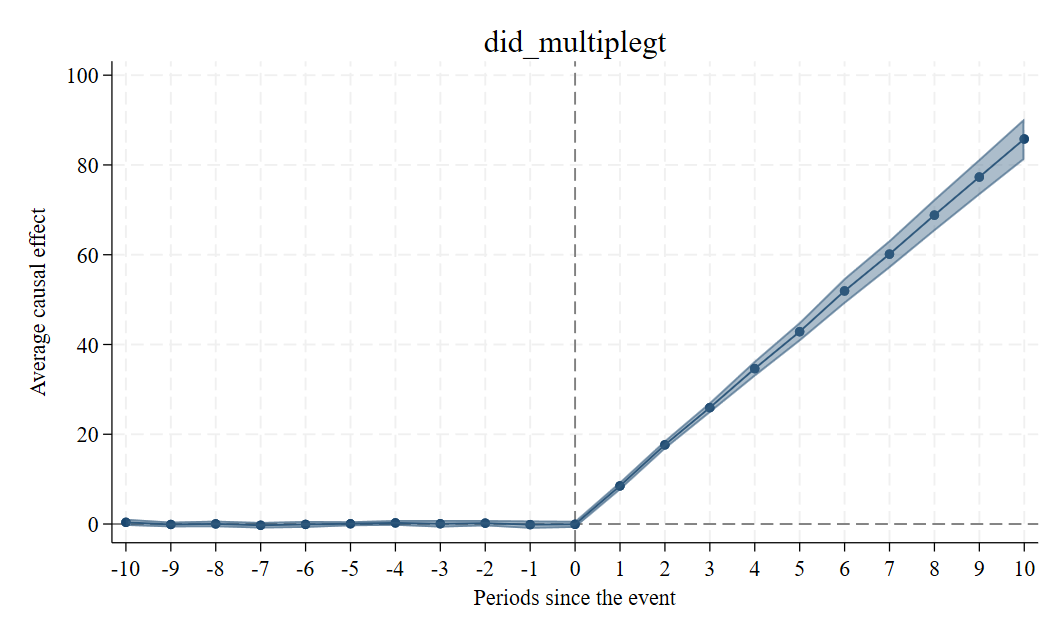
did_multiplegt (Chaisemartin and D’Haultfœuille 2020, 2021)
Note
To estimate event-study/dynamic effects, we strongly recommend using the much faster did_multiplegt_dyn command.
In addition to that, did_multiplegt_dyn offers more options than did_multiplegt, among which:
- normalized: estimation of the normalized dynamic effects (de Chaisemartin & D’Haultfoeuille, 2024);
- predict_het: built-in treatment effect heterogeneity analysis;
- design and date_first_switch: post-estimation options to analyze the design and timing of the treatment;
- by and by_path: estimating dynamic effects within levels of a group-level variable or within treatment paths;
- trends_lin: built-in group-specific linear trends.
Lastly, as of the last release, did_multiplegt_dyn also includes two user-requested features:
- only_never_switchers: restricting the estimators from de Chaisemartin & D’Haultfoeuille (2024) to only compare switchers and never-switchers;
- integration with esttab: here a quick tutorial.
Table of contents
Installation and options
ssc install did_multiplegt, replace
Take a look at the help file:
help did_multiplegt
Test the command
Please make sure that you generate the data using the script given here
Let’s try the basic did_multiplegt command:
did_multiplegt Y id t D, robust_dynamic dynamic(10) placebo(10) breps(20) cluster(id) seed(0)
and we get this output:
DID estimators of the instantaneous treatment effect, of dynamic treatment effects if the dynamic option is used,
and of placebo tests of the parallel trends assumption if the placebo option is used.
The estimators are robust to heterogeneous effects, and to dynamic effects if the robust_dynamic option is used.
| Estimate SE LB CI UB CI N Switchers
-------------+-----------------------------------------------------------------
Effect_0 | -.0608394 .3777101 -.8011512 .6794724 78 23
Effect_1 | 8.49767 .371304 7.769914 9.225426 78 23
Effect_2 | 17.64773 .4621524 16.74191 18.55355 78 23
Effect_3 | 25.9377 .6017678 24.75823 27.11716 78 23
Effect_4 | 34.62362 .9410695 32.77913 36.46812 75 23
Effect_5 | 42.85682 1.30207 40.30476 45.40888 64 19
Effect_6 | 51.93103 1.587562 48.81941 55.04265 64 19
Effect_7 | 60.13327 1.959996 56.29168 63.97486 64 19
Effect_8 | 68.82446 2.144383 64.62147 73.02745 64 19
Effect_9 | 77.30792 2.51719 72.37423 82.24161 64 19
Effect_10 | 85.78878 2.878414 80.14709 91.43047 55 19
Average | 40.79851 1.198362 38.44972 43.1473 762 229
Placebo_1 | .1308918 .5623093 -.9712345 1.233018 78 23
Placebo_2 | -.0635463 .3870331 -.8221312 .6950386 78 23
Placebo_3 | -.1275425 .4243438 -.9592563 .7041713 78 23
Placebo_4 | -.3848304 .3591436 -1.088752 .3190911 78 23
Placebo_5 | -.4583827 .294038 -1.034697 .1179317 75 23
Placebo_6 | -.1875761 .3982451 -.9681365 .5929843 64 19
Placebo_7 | .1194069 .4310281 -.7254083 .9642221 64 19
Placebo_8 | .0628537 .4707887 -.859892 .9855995 64 19
Placebo_9 | -.1943704 .4800074 -1.135185 .746444 64 19
Placebo_10 | -.2936845 .4425423 -1.161067 .5736984 64 19
The command also produces by default an event-study graph, unless the firstdiff_placebo option is specified: in that case, we do not recommend putting together first-difference placebos and long-difference event-study estimates on the same event-study graph.
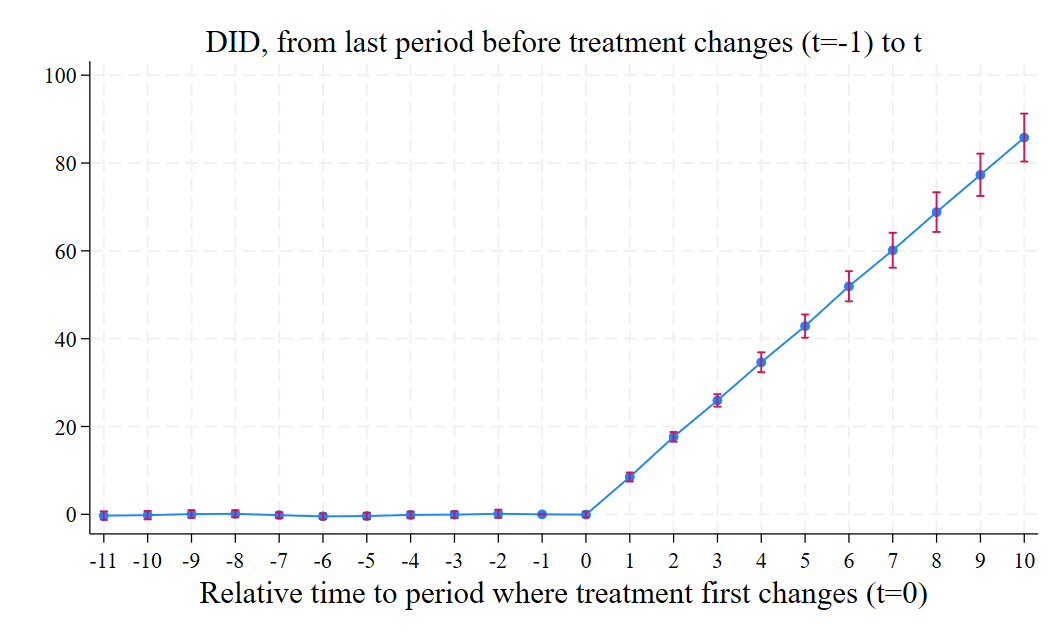
We can also plot the results using the event_plot (ssc install event_plot, replace) command as follows:
event_plot e(estimates)#e(variances), default_look ///
graph_opt(xtitle("Periods since the event") ytitle("Average causal effect") ///
title("did_multiplegt") xlabel(-10(1)10)) stub_lag(Effect_#) stub_lead(Placebo_#) together
and we get this figure: